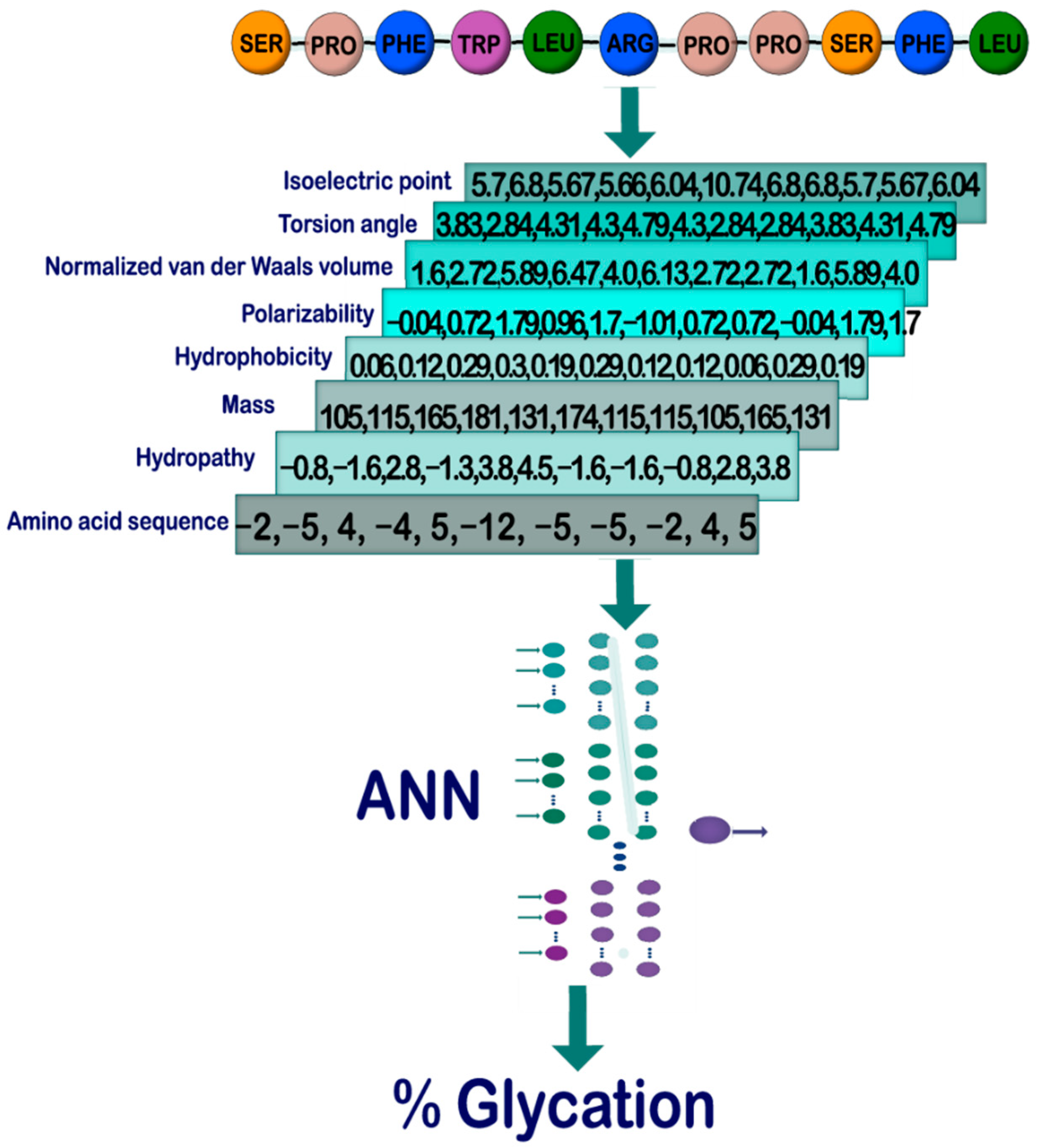
Pyteomics 4.0: Five Years of Development of a Python Proteomics Framework | Journal of Proteome Research
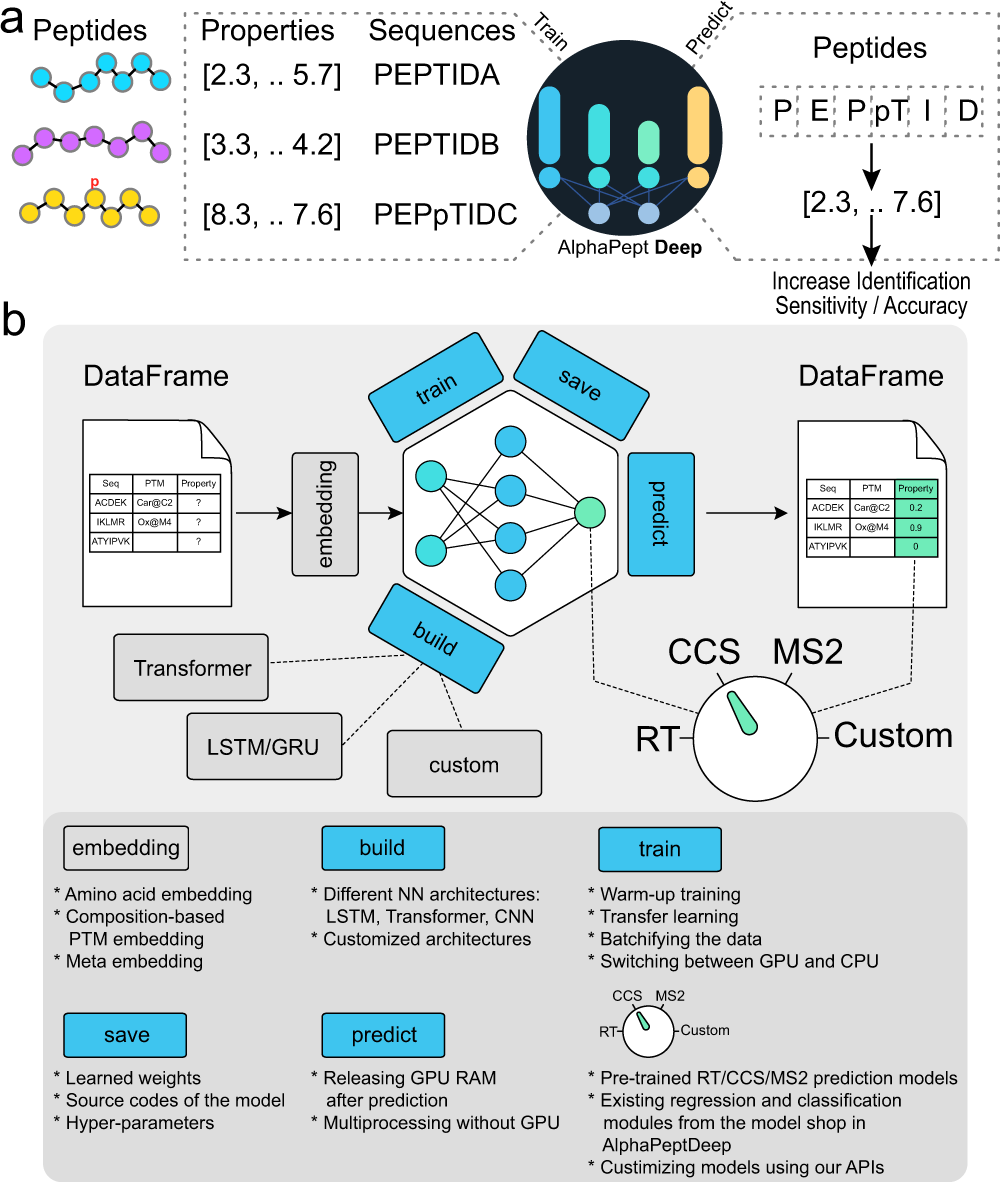
AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics | Nature Communications
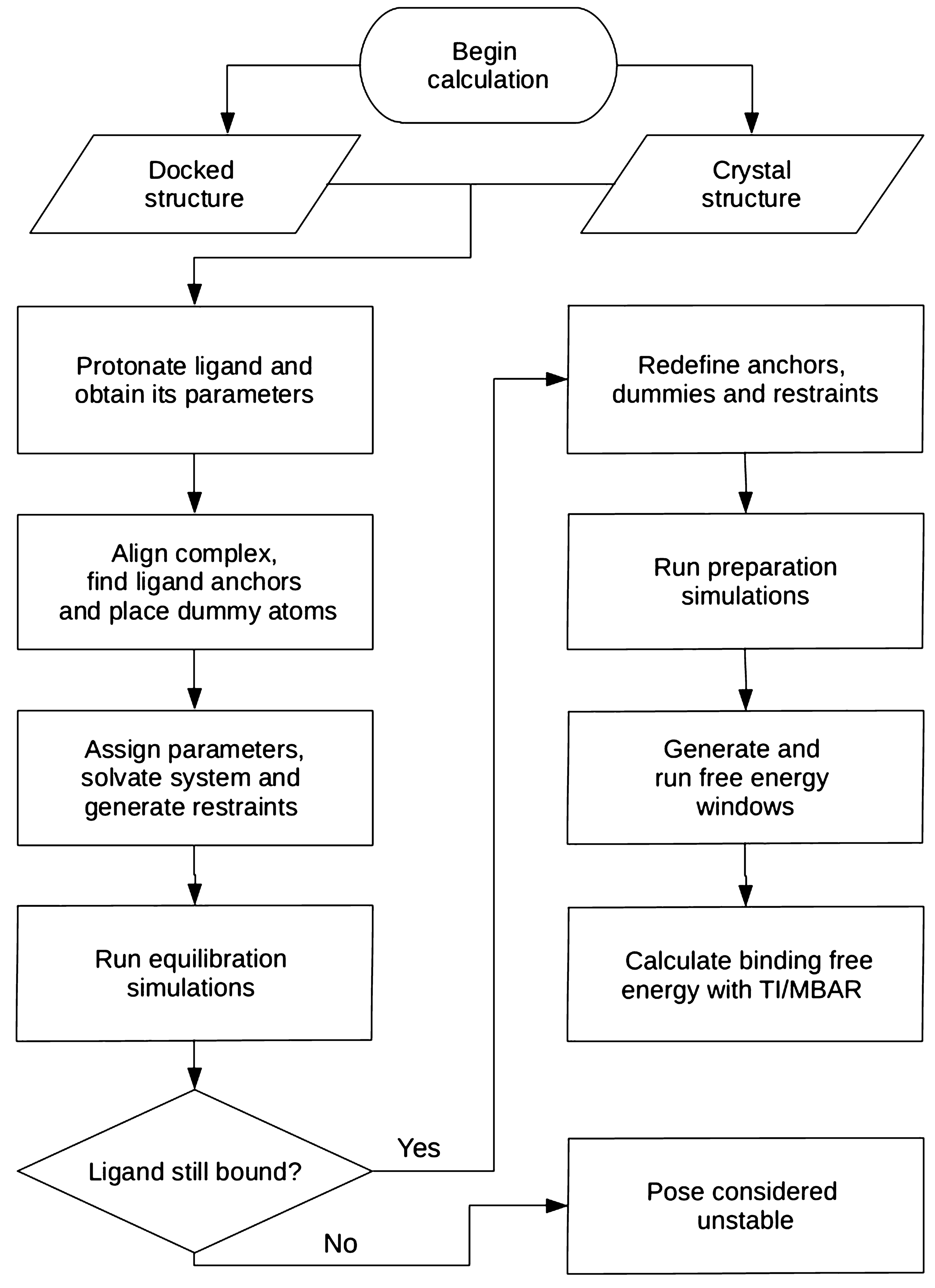
Automation of absolute protein-ligand binding free energy calculations for docking refinement and compound evaluation | Scientific Reports




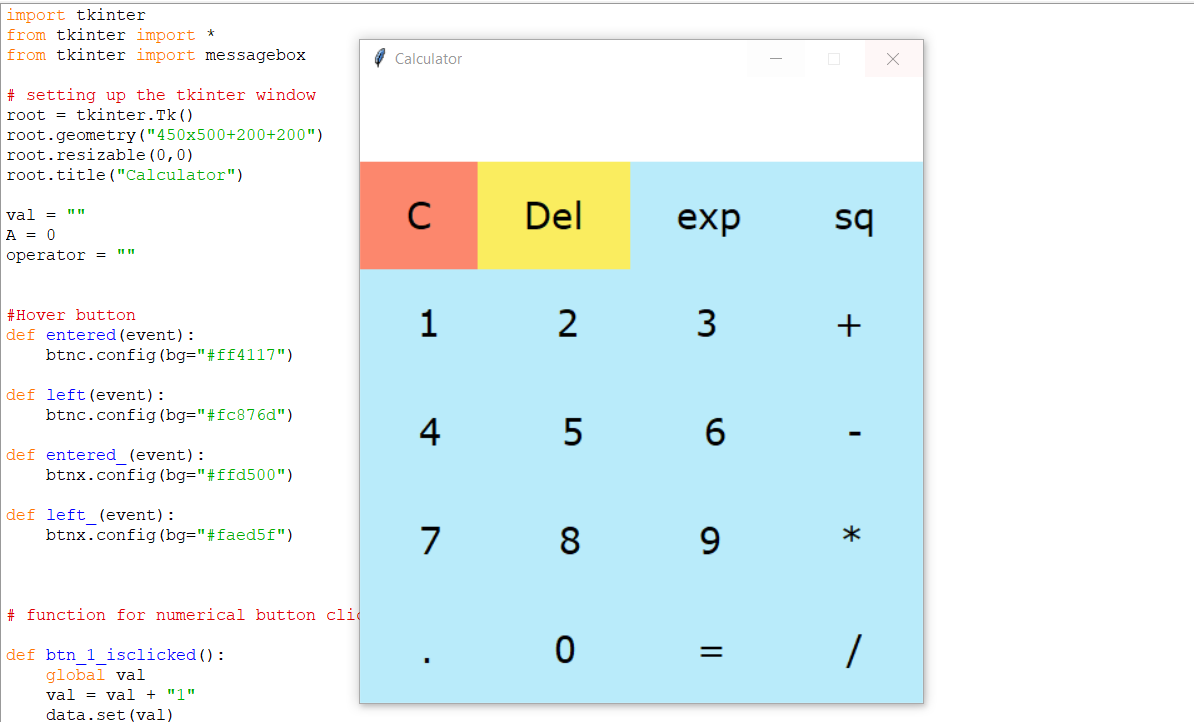
![Code for GUI Calculator in Python [Explained in Detail] Code for GUI Calculator in Python [Explained in Detail]](http://www.csestack.org/wp-content/uploads/2017/07/command-line-Python-Calculator.png)